Principal Component Analysis - Derived & Coded
Jul 7, 2023Principal Component Analysis (PCA) is a method to get low dimensional representation of a high-dimensional dataset without losing the important features. PCA can be useful for visualization of the dataset or determining features from data. More formally, if we are given data set of $n$ original $d$-dimensional points $X = \{x_1,\dots,x_n\} \in \mathbb{R}^{n\times d}$, we want to obtain $n$ new $k$-dimensional points $Z = \{z_1, \dots, z_n\} \in \mathbb{R}^{n\times k}$ with $k\ll d$, for which the projection back onto the original dimension $d$ ($\bar X = \{\bar x_1,\dots,\bar x_n\} \in \mathbb{R}^{n\times d}$), is as close as possible to the original data $X$.
 Nice way of visualizing the distance
Nice way of visualizing the distance between $\bar X$ and $X$ [builtin.com]
SVD decomposition of matrix $A$ is a representation of that matrix as $$A = USV^\top$$ where $U,V$ are orthogonal matrices and $S$ is diagonal matrix with singular values. We can (together with eigenvectors) reorder them such that $\lambda_1 \geq \lambda_2 \geq \dots \lambda_d \geq 0$. That will be important for the next step.
Hence the SVD decomposition goes as follows.
$$
n \Sigma={X}^T {X}={V} {S}^T \underbrace{{U}^T {U}}_{=I\ (\text{orthogonal})} {S} {V}^T={V} {S}^T {S} {V}^T={V} {D} {V}^T
$$
And so (assuming the decreasing order of the eigenvalues) the solution is given by $v_1$ for $k=1$ - the first column of $V = \begin{pmatrix} v_1 | \dots |v_d \end{pmatrix}$ from the SVD, the eigenvector corresponding to the biggest eigenvalue. This is the vector carrying the most amount of information about the dataset.
In case of PCA for $k\geq1$, the problem generalizes into
$$
(W^*,Z) = \arg\min_{W^\top W = I, Z} \sum_{i=1}^n \|Wz_i - x_i \|^2_2
$$
We can solve it analogously to $k=1$.
Obtaining $W$ - We can generalize the previous case for $k=1$ and say that the solution is given by $k$ eigenvectors corresponging the $k$ largest eigenvalues of the SVD decomposition on $X^\top X$, i.e. $$W = (v_1| \dots| v_k)$$
Obtaining $Z$ - As we already know how to get $z_i$ using $w$ from the $k=1$ case, it generalizes into $z_i = W^\top x_i$ for all $i=1,\dots ,n$, i.e. $$Z = W^\top \cdot X$$
- Standardize the data
- Compute the Covariance matrix of $X$
- Decompose this matrix to get $k$ eigenvectors corresponding to the $k$ highest eigenvalues
- Stack them vertically to get matrix $W = (v_1| \dots| v_k)$
- Project the original data to obtain $ Z = (z_1,\dots,z_n)$
From scratch implementation
To make this implementation demonstration a little more robust, we will no longer assume standardized data. I will demonstrate each part of the implementation on an example. Let's say we want to perform PCA with $k=1$ onto a dataset $X$Generating & Standardizing Dataset $X$
There are many datasets availbale for PC Analysis, but I thought it would be more fun to generate the data manually. The functiongenerate_points_line() takes 4 arguments. min_x, max_x determine the range of x-axis, deviation allows the point to be a little further away from the $y=x$ line and the 2-dimensional point will be taken to the dataset with probability p.
def generate_points_line(min_x, max_x, deviation, p):
points = list()
for x in range(min_x, max_x+1):
for y in range(min_x, max_x+1):
if y < x+deviation and y > x-deviation:
if (random.random() < p):
points.append((x,y))
return points
X = generate_points_line(0,120,50,0.23)Using the parameters min_x = 0, max_x = 120,
deviation = 50 and p = .023, I generated the following dataset $X$.
X = (X-np.mean(X, axis=0)) / np.std(X, axis=0) # standardize the data 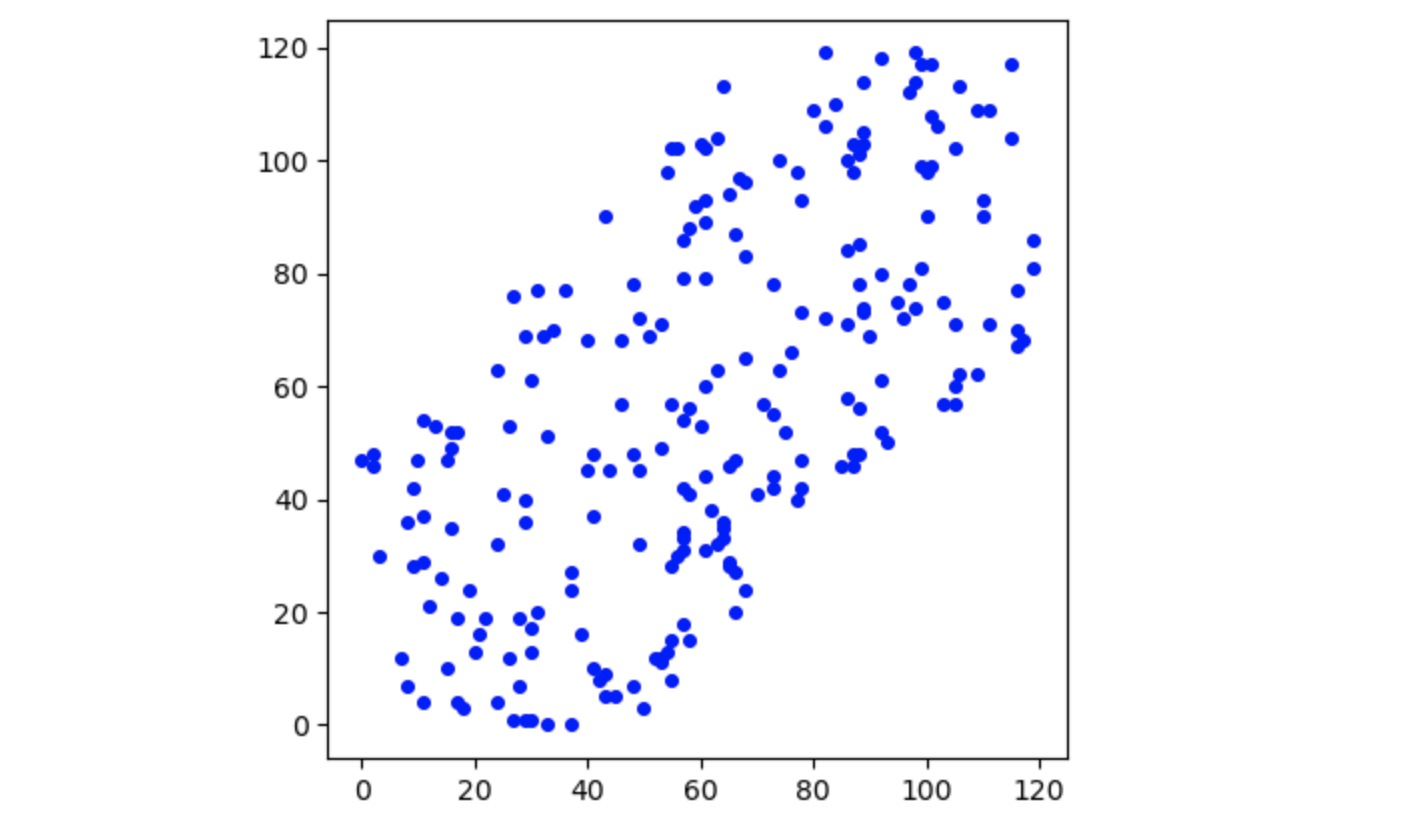 Generated Dataset
Generated Dataset
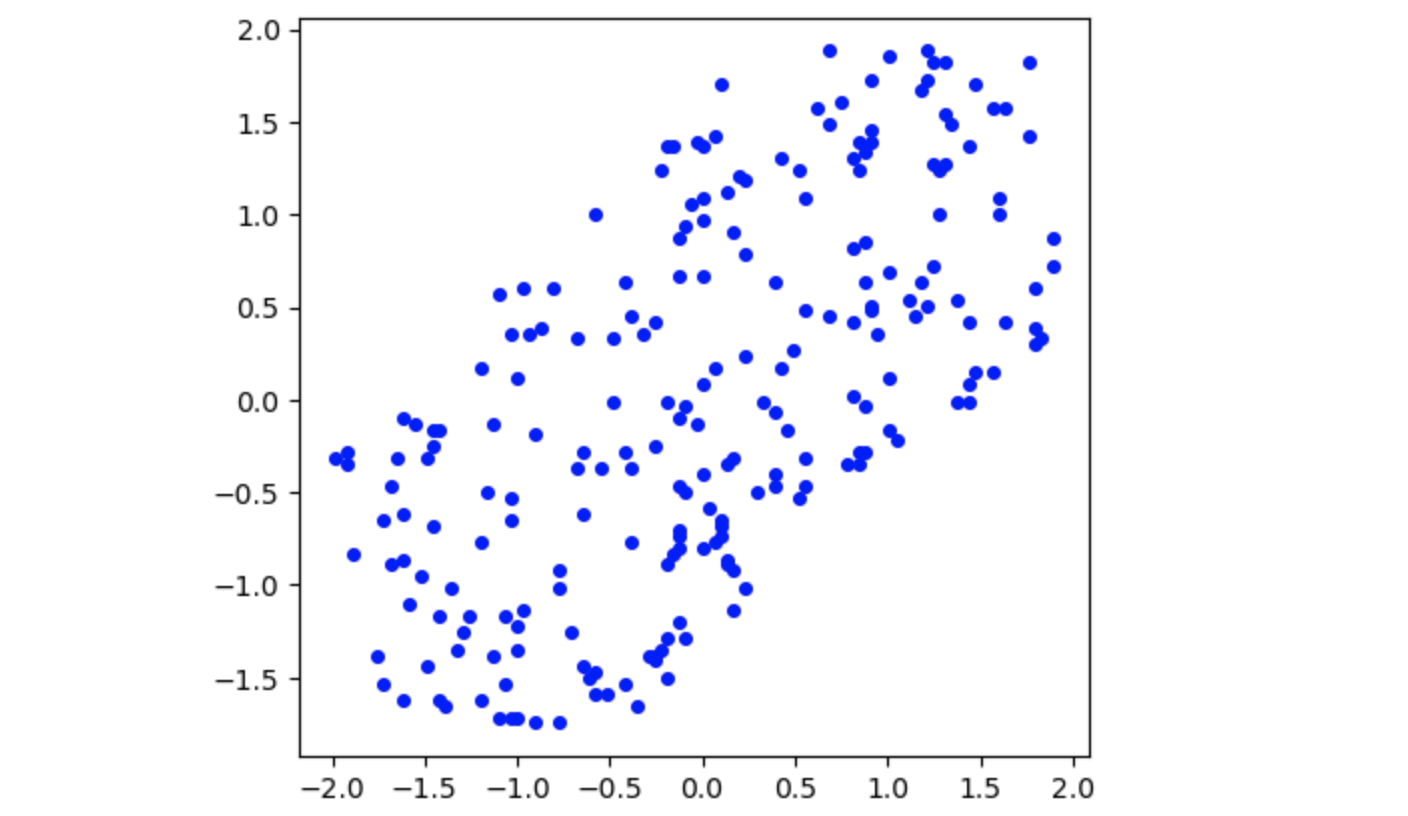 Standardized Dataset
Standardized Dataset
Covariance matrix
Computing the covariance matrix has now become easier since the $mean_X=0$ and $std_X=1$. All we need to do is $$ Cov = \frac{X^\top X}{n} \quad n=\text{number of points} $$def cov_of_standardized(X):
n = X.shape[0]
return np.dot(X.T,X)/(n)
cov_X = cov_of_standardized(X)This is not a suprising result as the covariance matrix of X is a square matrix where the diagonal elements are the variances of the variables. Since each standardized variable has a variance of 1, the diagonal elements of the covariance matrix of a standardized dataset are all 1.
Getting the Eigenvectors of the Covariance Matrix
Just for fun, let's see the eigenvalues (we expect two) and the corresponding eigenvectors (we expect 2x(1,2) vectors):U,s,Vt = np.linalg.svd(cov_X) # https://numpy.org/doc/stable/reference/generated/numpy.linalg.svd.html
print(s)
print(U)[179.23427235 41.76572765] # eigenvalues
[[-0.70710678 -0.70710678] # first eigenvector
[-0.70710678 0.70710678]] # second eigenvector[-0.70710678 0.70710678] component and the red one corresponds to the [-0.70710678 -0.70710678] component.
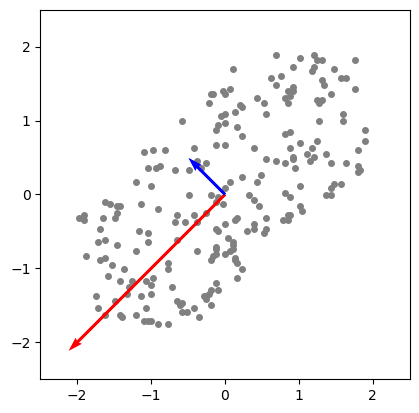 PCA Components. Eigenvectors of the $Cov$ matrix scaled by their respective eigenvalue
PCA Components. Eigenvectors of the $Cov$ matrix scaled by their respective eigenvalue
[-0.70710678, -0.70710678] to be extracted).
In the following snippet I implemented exactly that:
def k_most_eigenvectors(mat, k):
u,s,vt = np.linalg.svd(mat) # https://numpy.org/doc/stable/reference/generated/numpy.linalg.svd.html
indices = np.argsort(s) # returns indices from *smallest* to biggest EValue
indices = np.flip(indices,0)[:k] # (data, axis) and up to k-th index
W = np.real(u[:,indices]) # more context: https://numpy.org/doc/stable/reference/generated/numpy.linalg.svd.html
return W
W = k_most_eigenvectors(cov, 1) # extract the 1 most significant eigenvector of the covariance of X
W[-0.70710678, -0.70710678]Mapping the data
Last think we have to do is map the old points and obtain the points: $Z = W^\top X$. Also remember that $\bar X = W\cdot Z$. This is simply:
def fit_pca(X, W):
return np.dot(X,W)
Z = fit_pca(X,W)
X_bar = np.dot(W,Z.T)And now we're done! We can plot the new points $(z_1, \dots, z_n)$ in orange against the original points in blue to visualize $Z$. We can do the same also with the re-projected points $(\bar x_1, \dots, \bar x_n)$ (distance of which from original dataset we wanted to minimize) and get a nice visualization of the dimension reduction.
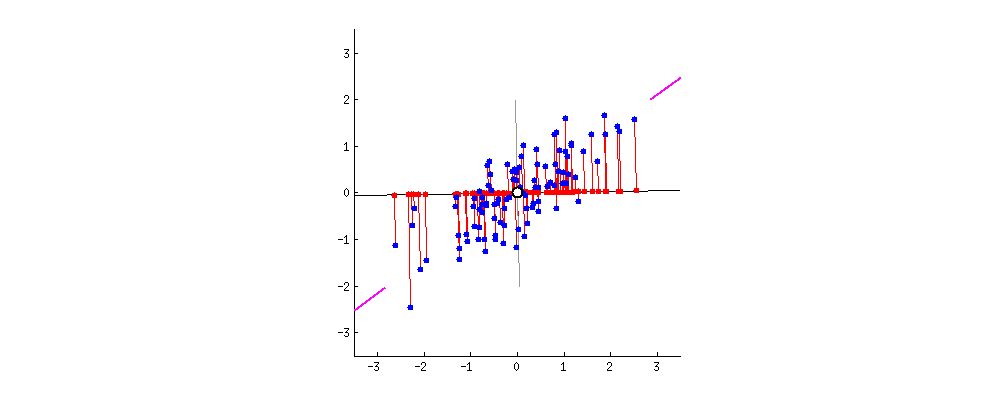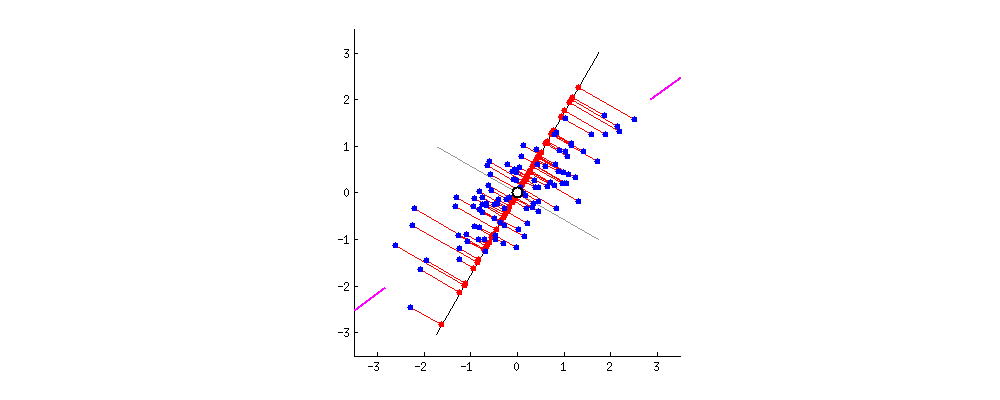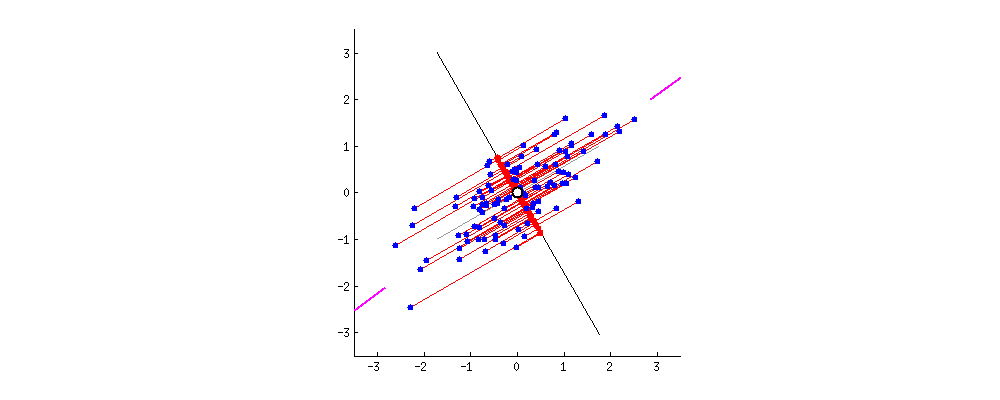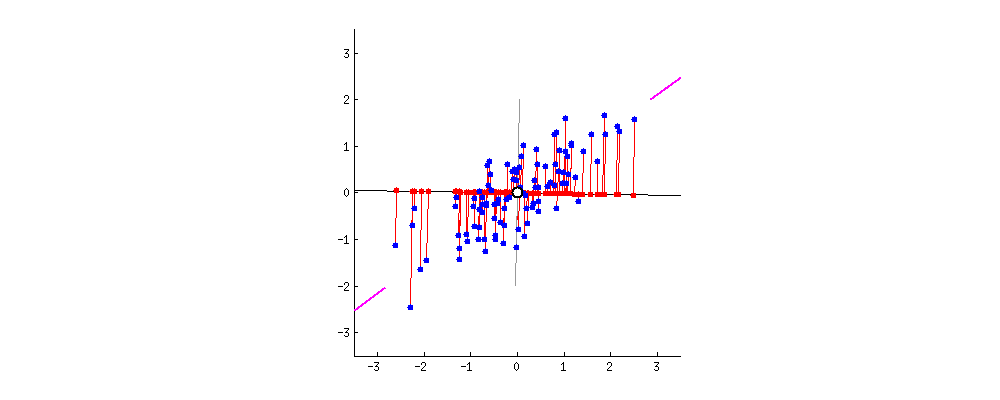
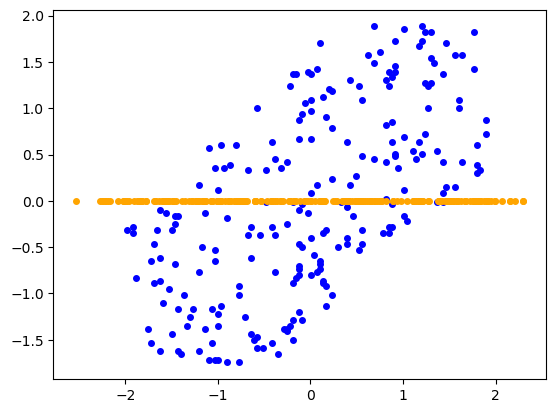 Obtained dataset $Z$ - PCA on $X$ with $k=1$
Obtained dataset $Z$ - PCA on $X$ with $k=1$
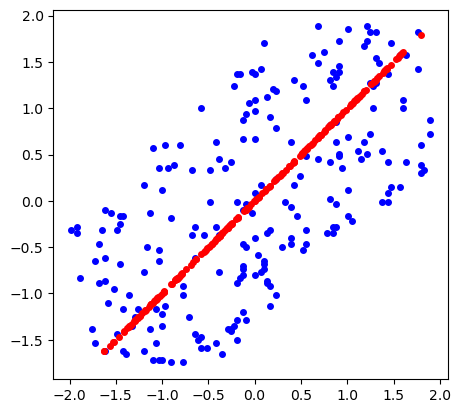 Retrieved points $\bar X$
Retrieved points $\bar X$
Important to notice - we are only considering the bigger eigenvalue for this dimension reduction. This means the larger eigenvalue is contains around $179.23427235 / (179.23427235 + 41\\.76572765) \approx 81\%$ of the information about the original dataset. This can be useful if we want for example to sustain given percentage of the original dataset features - we can choose $k$, such that the sum of the eigenvalues satisfies the percentage.